 The RAPIDS machine learning library, cuML, supports several types of input data formats while attempting to return results in the output format that fits best into users’ workflows. The RAPIDS team has added functionality to cuML to support diverse types of users: Maximize compatibility Users with existing NumPy, Scikit-learn, and traditional PyData libraries-based workflows: cuML’s … Continued
The RAPIDS machine learning library, cuML, supports several types of input data formats while attempting to return results in the output format that fits best into users’ workflows. The RAPIDS team has added functionality to cuML to support diverse types of users: Maximize compatibility Users with existing NumPy, Scikit-learn, and traditional PyData libraries-based workflows: cuML’s … Continued
The RAPIDS machine learning library, cuML, supports several types of input data formats while attempting to return results in the output format that fits best into users’ workflows. The RAPIDS team has added functionality to cuML to support diverse types of users:
Maximize compatibility
Users with existing NumPy, Scikit-learn, and traditional PyData libraries-based workflows: cuML’s default behavior, allowing as many formats as possible, and its Scikit-learn based API design, allows for porting parts of these workflows with minimal effort and no disruptions. So, for example, you can use NumPy arrays for input and get back NumPy arrays as output, exactly as you expect, just much faster.
Maximize performance
Users who want ultimate performance by keeping everything in the GPU’s memory: cuML’s use of open-source standards and configurability of behavior allows users to achieve maximum performance with low effort. This post will go into the details of how users can leverage this work to get the most benefits from cuML and GPUs.
Compatible input formats: the wonders of the CUDA Array Interface
Thanks in great part to the cuda_array_interface, referred to as CAI, cuML accepts a multitude of data formats:
- cuDF objects (DataFrames and Series)
- Pandas objects (DataFrames and Series)
- Numpy arrays
- CuPy and Numba device arrays
- Any CAI-compliant object, like PyTorch and CuPy arrays. This group is referred to as CAI arrays.
This list is constantly expanding based on user demand. For example, the cuML team is working on direct support for the dlpack array standard, coinciding nicely with TensorFlow’s new support. This can also be done by going through either cuDF or CuPy, which also have dlpacksupport. If you have a specific data format that is not currently supported, please submit an issue or pull request on Github.
Default Behavior: how does cuML work out of the box?
cuML’s default behavior is designed to mirror the input as much as possible. So, for example, if you are doing your ETL in cuDF, which is very typical for RAPIDS users, you would see something like:
When you use cuDF DataFrames, cuML gives you back cuDF objects (in this case, a Series) as a result. But, as mentioned preceding, cuML also allows you to use NumPy arrays without changing the cuML call:
In this case, now cuML gives back the results as NumPy arrays. Mirroring the input data type format is the default behavior of cuML, and in general, the behavior is:
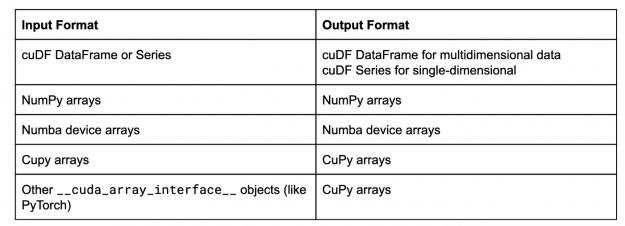
This list is constantly growing, so expect to see things like dlpack compatible libraries in that table soon.
Configurability: how do I make cuML behave my way?
cuML allows users to configure output types globally. For example, if your ETL and Machine Learning workflow is GPU-based, but you rely on a NumPy based visualization framework, try this:
Using the set_global_output_type instruction affects all subsequent calls to cuML. In case users want finer-grained control (for example, your models are processed by GPU libraries, but only one model needs to be NumPy arrays for your specialized visualization), the following mechanisms are available:
- cuML’s context manager
using_output_type:
- Setting the output type of individual models:
This new functionality automatically converts data into convenient formats without manual data conversion from multiple types. Here are the rules that the models follow to understand what to return:
- If output_type was specified when building the model, for example
cuml.KMeans(n_clusters=2, output_type=’numpy’), then it will give results in that type. - If the model was built inside a context manager
withusingcuml.using_output_type, then the model uses the output_type of that context. - If the
output_typewas set usingset_global_output_type, then it will give back that type of results. - If none of the preceding are specified, then the model will mirror the type of the objects used for input, as described in the default behavior section.
Efficiency: what formats should I use?
Now that you know how to use cuML’s input and output configurability, the question is, what are the best formats to use? It will depend on your needs and priorities since all formats have trade-offs. Let’s consider a simple workflow:

Using NumPy based objects
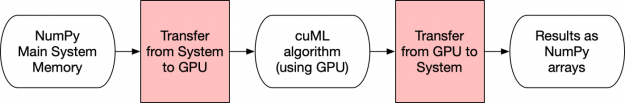
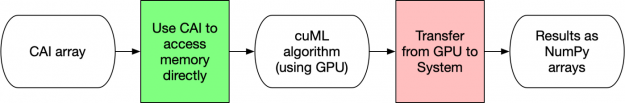
In Figure 3 below, the transfers (pink boxes) limit the amount of speedup that cuML can give you since the communications use the slower system memory and you have to go through the PCI Express bus. Every time you use a NumPy array as input to a model or ask a model to give you back NumPy arrays, there is at least one memory transfer between the main system memory and the GPU.
At first glance, one might imagine that doesn’t impact much. Yet keeping data as much as possible in the GPU is one of the, if not the biggest reason, RAPIDS achieves its lightning speed.
Using cuDF objects
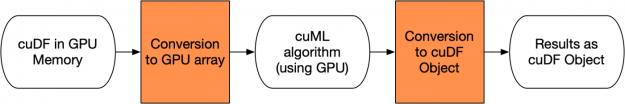
Using GPU objects as opposed to NumPy arrays has significant implications. For example, using cuDF objects is illustrated in Figure 4 below. The orange boxes represent conversions that happen entirely on the fast GPU memory. Unfortunately, this means that an extra copy of the data will be done during the cuML algorithm processing, which can limit the size of the dataset that can be processed in a particular GPU.
DataFrames (and Series) are very powerful objects that allow users to do ETL in an approachable and familiar manner. But to offer this, they are complex structures with significant amounts of complexity to enable this functionality.
A few examples of this are:
- Every column can have, besides its data, a bitmask array (essentially an added array of zeros and ones) that allows users to have missing entries in their data.
- Due to the flexibility that DataFrames need to provide for adding/removing rows and columns, each column might be far away from each other in memory.
- And of course, there are added structures for things like indexes and column names.
However, these constraints present some difficulties for some analytics workflows:
- First, many algorithms work significantly better when all your data is contiguous, for example, all the bytes are grouped together in the same memory region, since accessing memory efficiently is a huge component of processing data fast (particularly for GPUs!).
- Memory is a limited resource (in general, but even more so for GPUs and accelerators), so the added overheads can have a very significant impact.
Using device arrays
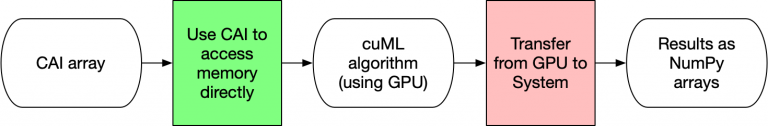
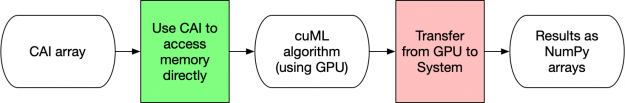
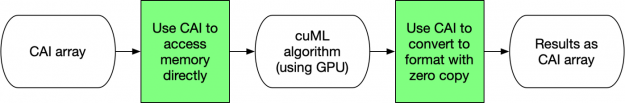
Figure 5 below illustrates how CAI arrays for input or output have the lowest overhead for processing data in cuML. By using the CAI, no memory transfers nor conversions occur. cuML uses the attributes of the CAI directly to access the data and then return a CAI array. There is virtually no overhead for these formats. Device arrays, such as those from CuPy or Numba, are significantly simpler structures than the DataFrame/Series equivalents. Similar to NumPy, they are designed to be contiguous blocks of memory that are described by metadata. This design decision is why NumPy was revolutionary for the original Python ecosystem. Given all of this, it shouldn’t be a surprise that device arrays are the most efficient way of using cuML!
As mentioned preceding, all CAI arrays are essentially the same from cuML’s perspective, so your workflows could combine functions of Numba, CuPy, cuML, and so on without needing to do expensive memory copying operations.
Tips for selecting data types
So what data type should you use? As mentioned before, it depends on the scenario, but here are a few suggestions:
- f you have an existing PyData workflow, take advantage of cuML’s NumPy functionality to try different models piece by piece. Start by accelerating the slowest parts of your workflows. DBSCAN and UMAP are great examples of modInels in cuML that even when used by themselves, without full RAPIDS acceleration, provide huge speedups and improvements.
- Potential pitfall: This could create a communication bottleneck between the main system memory and the GPU memory.
- If your workflow is very ETL-heavy with lots and lots of cuDF work, where the bulk of the processing and development time is in data loading or transformation, keep things as cuDF objects and let cuML manage conversions.
- Potential pitfall: This might limit how much data you can fit for a single model in a GPU.
- If ultimate speed of training or inference is the key part, then adapt your workflow to use CUDArray interface libraries as much as possible.
With all of these tips, you can configure cuML to optimize your needs as well as better estimate the impacts and bottlenecks of workflows. Your new workflow may now look something like this:
What’s next?
Here are some active areas we are excited to share in upcoming posts:
Multi-Node Multi-GPU (MNMG) cuML: There is much additional work being done. Many engineers on the RAPIDS cuML team are currently building Multi-Node Multi-GPU (MNMG) implementations of leading algorithms to enable distributed machine learning at scale. Distributed data is an entire topic by itself, with more posts coming soon. But as of version 0.13, MNMG cuML accepts Dask-cuDF objects (the distributed equivalent of cuDF using Dask) and CuPy backed Dask Arrays. cuML produces results in MNMG algorithms that mirror the input you use, similar to the default behavior of cuML for a single GPU. We are working on adding more configurability to the MNMG cuML algorithms. We will talk about how your data is distributed, and what formats you use, impact cuML.
Lower-level details about your data and its implications: Many details, like datatypes or the ordering of the data in memory, can affect cuML. We will talk about how those details affect cuML, and how it compares and differs from traditional PyData libraries.
Abstractions and design: Recently introduced abstractions and mechanisms in the RAPIDS software stack, like the CumlArray, allow cuML to provide this functionality while reducing code complexity and the number of tests needed to guarantee results. We will talk about how this, alongside the CAI, gives users the ability to use multiple libraries like CuPy, cuDF, cuML together with very little effort.
Conclusion
This post discussed the input and output configurability capabilities of cuML, the different data formats supported, and the advantages and disadvantages of each format in cuML. The post shows how easy it is to adopt cuML into existing workflows. cuML’s scikit-learn API and output mirroring of formats allow you to use it as a drop-in replacement for existing libraries. To extract the maximum performance, users should try using GPU-specific formats as much as possible, and CAI arrays like CuPy or Numba. The RAPIDS team is working on improving cuML’s capabilities and supported data formats. If you have an interest in some particular format or some functionality that would improve cuML for your use-cases, raise an issue in the cuML Github repository, or come chat with the team in the RAPIDS slack channel.
